SOFTWARE FOR MATERIALS MODELING AND SIMULATION
The multidisciplinary and integrated modeling approach often pursued in industrial modeling and simulation was introduced, and the multiscale modeling paradigm usually preferred for such work was summarized, in a post titled MULTISCALE MODELING AND SIMULATION IN INDUSTRY (created 17 November 2018).
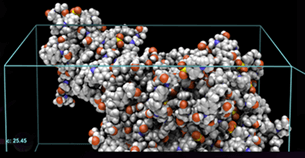
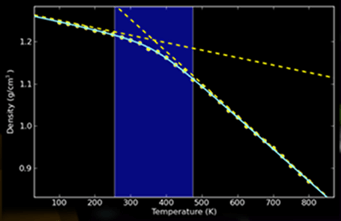
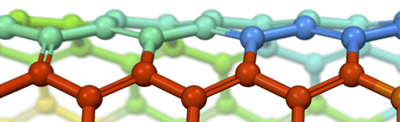
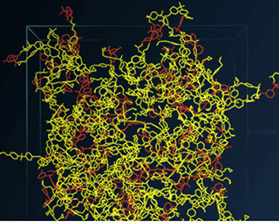
The present post takes the discussion one step further by presenting overviews of the materials modeling and simulation software programs that are commercially available from four leading developers of such software. The images shown in each section are reproduced from the product literature of the software developer that is being discussed in that section.
The choices of best supplier and of optimum combination of software capabilities depend on the needs of each organization and must be considered carefully to make the best decisions. Since there is no general answer to the question of which software developer offers the most useful products, the software developers will be discussed below in alphabetical order.
Dr. Bicerano has 45 years of experience with materials modeling and simulation. During his consulting practice, his work has included providing strategic guidance to clients in planning the materials modeling and simulation component of their R&D program, helping clients select the best methods and software for their needs, and solving specific problems.
MATERIALS DESIGN, INC.
Materials Design, Inc. provides a broad range of software tools for materials modeling and simulation integrated into the MedeA environment. MedeA enables professional, day-to-day deployment of atomic-scale and nanoscale computations for materials engineering, materials optimization, and materials discovery. Powerful simulation engines are integrated in MedeA with elaborate property prediction modules, experimental databases, structure builders, and analysis tools, all in one user-friendly environment. MedeA supports chemical, metallurgical, electronic, polymeric, and materials science research applications.
The capabilities and benefits of the MedeA environment are summarized below. These capabilities include Polymer Expert, a polymer informatics software tool for de novo polymer design (please watch this introductory webinar) that Dr. Bicerano developed in collaboration with Materials Design, Inc.

As shown below, all classes of materials of industrial interest can be studied on the MedeA environment, and a broad range of properties can be calculated for these materials.

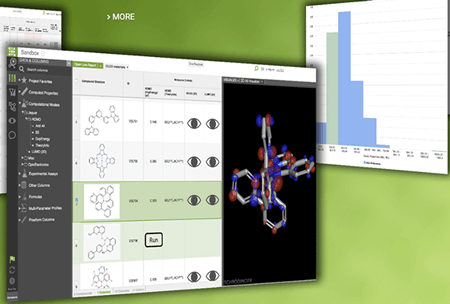
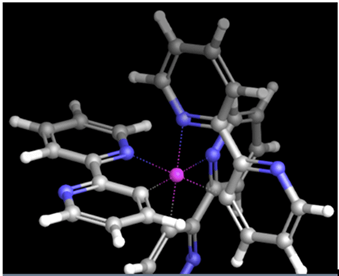
The following are examples of what a user may see on the screen while working on a problem on the MedeA environment.

MedeA's layered architecture connects workflows with modern computing environments to create a highly productive environment. MedeA has a 3-tier architecture that offers flexibility, and optimizes the use of available computing resources, while reducing the human time and effort involved in simulation deployment. This 3-tier architecture consists of the MedeA Environment (a graphical use interface that is the main tier), JobServer (middle tier), and TaskServer (end tier).
The user interacts with the MedeA Environment and submits a Job (a computational protocol that applies to one or more structures) to the JobServer. A Job can require multiple simulation runs of specific compute engines; such as VASP, LAMMPS, GAUSSIAN, MOPAC, and GIBBS. Each of these runs is called a Task, and will be submitted to computing resources hosting TaskServers.
This 3-tier architecture of MedeA is shown below. The software tools integrated into each tier are also summarized in this image. Readers are referred to the company’s website for detailed information on each software tool.

In addition to the many software tools that are already built into it, MedeA also allows users to add other computation engines, databases, and CPUs, and thus expand their capabilities.
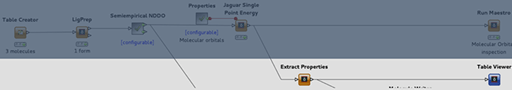
The following image further illustrates the three tiers of MedeA in the simplest case where a single graphical user interface (GUI) is connected to a single JobServer which is in turn connected to a single TaskServer. More complex implementations of this architecture, with multiple devices being interconnected at one, two, or all three tiers, are often used.

As summarized by the schematic depiction reproduced below, putting elaborate compute protocols into action is simple when using MedeA’s flowcharts which contain full information about the user’s computing jobs. Furthermore, since these flowcharts are stored by MedeA’s JobServer, the user can reload, share, and refine them over time.

High-throughput (HT) calculations are essential in today’s materials modeling practice. The MedeA HT-Launchpad and MedeA HT-Descriptors modules provide powerful and yet user-friendly tools to accomplish HT modeling tasks. These modules enable the user to generate large and consistent sets of computed data and to create descriptors combining experimental and computed properties. The user is thereby able to screen, understand, and optimize materials, thus also creating the input for machine learning procedures.

MedeA Instrument enables the user to skip the data center and gain more direct control of high-performance computing and simulation activities.

Links to many detailed examples of specific applications of the software are provided under the heading of All Application Notes.
BIOVIA
BIOVIA, which was previously named Accelrys, is a wholly owned subsidiary of Dassault Systèmes. It develops and sells a vast range of modeling and simulation software. The powerful BIOVIA product portfolio is focused on integrating the diversity of science, experimental processes, and information requirements across research, development, QA/QC, and manufacturing. Capabilities cover scientific data management; small molecule, biological, and materials modeling and simulation; cheminformatics and bioinformatics; systems biology and integrative therapeutics; collaborative network research; scientific pipelining; enterprise laboratory management; regulatory and quality management; process knowledge and collaboration; and chemical inventory management.
The remainder of the discussion of BIOVIA in this post will focus on its materials modeling and simulation software suite, named Materials Studio. This suite helps its users solve key materials and chemical research problems, within an integrated, multiscale modeling environment that delivers a complete range of simulation methods.
Materials Studio provides a complete modeling and simulation environment designed to allow researchers in materials science and chemistry to predict and understand the relationships of a material’s atomic and molecular structure with its properties and behavior. Materials Studio is used by researchers in many industries. The better-performing materials being developed with the help of Materials Studio include pharmaceuticals, catalysts, polymers, composites, metals, alloys, battery materials, and fuel cell materials. Materials Studio has helped companies accelerate innovation, reduce costs, improve efficiency, facilitate collaboration, and solve some of the most challenging materials science problems.
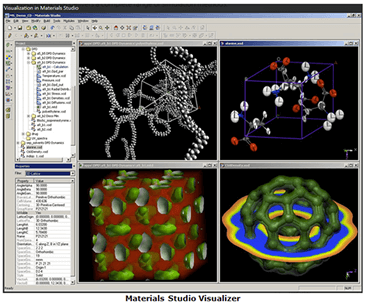
Materials Studio includes a graphical user environment (Materials Studio Visualizer) in which researchers can construct, manipulate, and view models of molecules, crystalline materials, surfaces, polymers, and mesoscale structures. Materials Studio Visualizer is complemented by a complete set of solution methods. These solution methods include quantum mechanical, atomistic (or “classical”), mesoscale, and statistical methods, thus enabling researchers to evaluate materials at various particle sizes and time scales. Materials Studio also includes tools for evaluating crystal structure and crystal growth.
Materials Studio is integrated with BIOVIA’s open, scalable, and scientifically aware Pipeline Pilot scientific informatics platform. Pipeline Pilot helps researchers by automating routine calculations, simplifying multistep workflows, and deploying routine calculations in web-based applications.
Expert scientists can encapsulate and automate best practices in reusable “protocols” using Pipeline Pilot and its scientific collections. These scientific collections include the Materials Studio Collection as well as the Polymer Properties (SYNTHIA) Collection based on J. Bicerano, Prediction of Polymer Properties, third edition, Marcel Dekker, New York (2002).

Materials Studio can also be integrated with third party or in-house applications to extend the breadth and depth of the science that it supports.

The many modeling and simulation tools available through Materials Studio are presented on the BIOVIA website in the following four groups:
- Analytical and Crystallization Software
![Software for materials modeling and simulation Software for materials modeling and simulation]()
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Polymers and Classical Simulation Software
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Quantum and Catalysis Software
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Visualization and Statistics Software (an image illustrating the capabilities of this software was reproduced earlier in this section of this post)
Many case studies related to the use of Materials Studio for materials science research are also presented.
CULGI
CULGI software includes all relevant aspects of computational chemistry. It scope ranges from quantum mechanical to molecular and coarse-grained modeling, to informatics and thermodynamics. A compelling feature of CULGI software is the concept of scripted workflows which is the key to its success in modeling a wide variety of systems.

CULGI software includes various empirical, semi-empirical, and ab-initio quantum mechanical calculation algorithms. These algorithms can be used to model the quantum mechanical behavior of solids (plane wave) and molecules. The range of available density functional methods is comprehensive and covers all popular functionals.

CULGI has an extensive suite of molecular dynamics algorithms, including a 3D molecular sketcher, a powerful atom-typer, and an amorphous builder. CULGI also has the capability to find new torsion force field parameters automatically. CULGI can read and write all popular molecular modeling formats. Even entire simulations can be imported and exported. The multiscale approach is also instrumental in building original structures. For example, in a typical scenario, one would first establish a coarse-grained polymer system. After relaxation and subsequent reverse mapping (coarse-grained to molecular), one would continue in molecular dynamics. Very complex initial structures can be created in this way.

CULGI is a leader in providing modeling solutions for nanostructured and microstructured chemical systems, whether from a synthetic or a biological origin. The principal method is a set of proprietary algorithms that enable the automatic coarse-graining of any chemical into fragments. The method is referred to as Automated Fragmentation and Parameterization. Coarse-grained simulation engines include Brownian and dissipative particle dynamics. The interaction models are comprehensive. They can consist of actives, charged systems, colloids, polymers, surfactants, and reactive systems. Coarse- grained force fields can be simple linear (as in dissipative particle dynamics), or more complex (by using tabular input).

CULGI has cutting-edge methods for mesoscopic dynamics simulations of complex polymer systems. These so-called dynamic density functional methods (or generalized phase-field models) allow the efficient calculation of submicron structures in composites, blends, and formulations.

CULGI has an extensive database of pre-calculated molecules and all useful descriptors known to date. This database serves as input for industrial projects. It is also used to calibrate the chemical informatics algorithms. CULGI includes a range of statistical calculation methods, such as linear and non-linear regression, and genetic algorithms. One of many ways in which one could extend the modeling capabilities is the interfacing of CULGI to machine learning software.

CULGI includes a recoded and reoptimized version of the powerful COSMO-RS methods for predicting the thermodynamic properties, such as pK, logP, and mixing Gibbs energies, from the molecular composition. COSMO-RS is particularly useful for the calibration of coarse-grained simulations. Another special feature of CULGI’s COSMO-RS is that it can offer the method based on superfast empirical and semi-empirical quantum mechanical calculations.

CULGI also includes the UNIFAC method for chemical engineering thermodynamics. This method is used to calculate various thermodynamic properties of industrial solvent mixtures, such as their Gibbs free energies of mixing. UNIFAC calculations are very rapid. The calculation of an activity coefficient typically takes less than one millisecond per system.

Robust mapping between different levels of modeling (quantum mechanical, molecular, coarse-grained, mesoscopic, and macroscopic) is instrumental in an industrial environment. CULGI includes methods for all kinds of mapping; from quantum mechanical to molecular, from molecular to coarse-grained and back, from coarse-grained to mesoscopic, etc.

The CULGI software can be used to solve problems in industries as diverse as oil and gas, chemicals, personal and home care, and pharmaceuticals. Examples of applications of the software in each of these industries are provided on the CULGI website. A white paper providing further details and additional case studies is available from CULGI upon request.
SCHRÖDINGER
Schrödinger provides the following five software suites:
- Materials science suite, providing an integrated solution for the atomic-scale simulation of chemical systems. It is used to compute the structures, reactivities, and properties of chemical systems, and thus advance the development of new materials and processes. This is the only suite that is highly relevant to the needs of a materials R&D environment where the emphasis is not on biological materials and/or drug development.
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Facilitation of all steps in an atomic-scale simulation project.
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Atomistic model builders and property workflows.
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Predictive capabilities across a wide range of chemical systems.
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Materials in silicodiscovery and optimization through automated simulation.
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- LiveDesign for Materials Science (a transformative collaboration, informatics, and machine learning platform).
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Maestro (a powerful, unified portal to modeling materials, small-molecules, and biologics with Schrödinger technology).
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- State-of-the-art visualization and workflow automation.
![Software for materials modeling and simulation Software for materials modeling and simulation]()
- Facilitation of all steps in an atomic-scale simulation project.
- Small-molecule drug discovery suite, to accelerate lead discovery and optimization.
- Biologics suite, to model biologics, antibodies, and proteins.
- Discovery informatics suite, which allows for data sharing and collaborative drug design across the entire discovery team in real time.
- PyMOL, providing high-performance molecular graphics for visualizing and communicating structural results. This suite can be beneficial in any R&D environment where there is a frequent need for molecular graphics of exceptional quality.
The components of the materials science suite are shown below.